Applied Bioinformatics
The applied bioinformatics task group within the International Lipidomics Society aims at facilitating the development of bioinformatics tools and workflows to foster the exchange of quantitative and qualitative lipidomics data.
This will be achieved by providing standardized data formats for data exchange between vendor software, the research community, bioinformatics applications, and public data repositories. Moreover, we aim at providing curated, high-quality datasets to facilitate the development and comparison of competitive and state-of-the-art bioinformatics algorithms and tools for lipid identification and quantification.
These goals are only achievable by continuing and intensifying the existing efforts for standardization of lipid nomenclature, data exchange formats, primarily, but not exclusively for lipidomics based on mass spectrometry and the curation and maintenance of high-quality reference databases for quantitative lipidomics, as well as structural and functional ontologies thereof.
By reaching these goals, we ensure better integration with other “omics” for more holistic and dynamic views onto the functions and roles of lipid metabolism in the context of systems biology and personalized medicine.
However, others have paved the way before us and have drawn the scientific community’s attention to the necessity of standardization of and FAIR access to primary and secondary research data. We thus also collaborate closely with existing societies, such as the HUPO-PSI and the Metabolomics Society to ensure alignment on common goals and to foster a combined momentum in moving the field forwards.
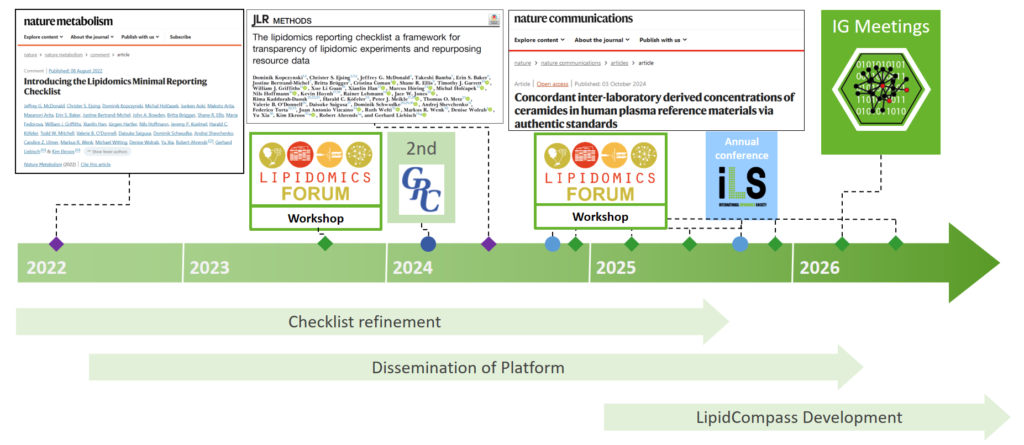
LipidCompass Dataset Curation
- Studies: 78
- Datasets: 165
- Biological Species: 16
- Tissues: 40
- Cell Types: 14
- Diseases: 0
Steering committee:
Dominik Kopczynski, Austria
Kevin Huynh, Australia
Jürgen Hartler, Austria
Nina Troppmaier, Austria
Bo Burla, Singapore
Checking out the Boundaries: Milestone in Lipidomics Achieved
Ring trial enables establishment of ceramide reference values Espoo, Finland – [Oct. 3rd, 2024] – Results of the first phase of a Ceramide Ring Trial
ELM 2020 Bioinformatics for Lipidomics Symposium
The ILS Applied Bioinformatics Interest Group invites you to join the Bioinformatics for Lipidomics Symposium on October 2nd, 2020, as part of this year’s joint
